X-RAY RUNS: Apply for Beamtime
2017 Nov 1 - Dec 21
2018 Feb 7 - Apr 3
2018 Proposal/BTR deadline: 12/1/17
2018 Apr 11 - Jun 4
2018 Proposal/BTR deadline: 2/1/18
Drugs which inhibit the activity of the reverse transcriptase of HIV-1 (HIV-RT) are effective in treating AIDS, since the virus cannot replicate without a working RT. Unfortunately, HIV is prone to mutations, some of which can render a formerly potent drug ineffective. The Arnold group (Rutgers) has now elucidated the mechanism by which a mutated form of HIV-RT resists the drug AZT, as reported in a recent publication:
Structural basis of HIV-1 resistance to AZT by excision
X. Tu, K. Das, Q. Han, J. Bauman, A. Clark Jr., X. Hou, Y. Frenkel, B. Gaffney, R. Jones, P. Boyer, S. Hughes, S. Sarafianos, and E. Arnold
Nature Structural & Molecular Biology (2010), Vol.
17, 1202-1209
AZT inhibits the DNA-to-RNA transcription carried out by HIV-RT by binding to and blocking the end of the DNA primer. HIV-RT is able to remove the AZT, using the cellular energy source ATP; this process is usually quite inefficient, but mutations to HIV-RT can enhance it to the point that AZT becomes ineffective. The Arnold group examined the crystal structure of HIV-RT with a suite of 5 such mutations, alone and in various complexes with DNA and AZT. They showed that the mutant had a high-affinity ATP-binding site not present in the wild-type protein, and concluded that the excision of AZT was promoted by binding of ATP to this new site.
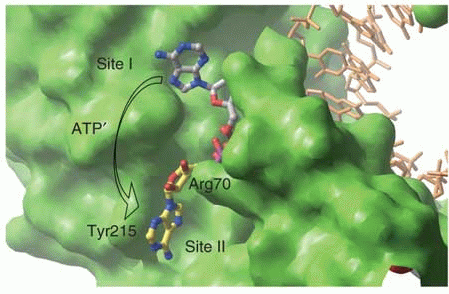
Binding sites of ATP on the surface of HIV-RT. Site I exists in both wild-type and mutant protein, but site II is present only in the mutant.
The crystal structures also helped to elucidate the mechanism by which HIV-RT transfers AZT from the 3' end of a DNA primer to ATP, hence unblocking the DNA, and suggested that drugs targeting the ATP binding site would inhibit this excision process and would be useful therapeutically when administered together with AZT.
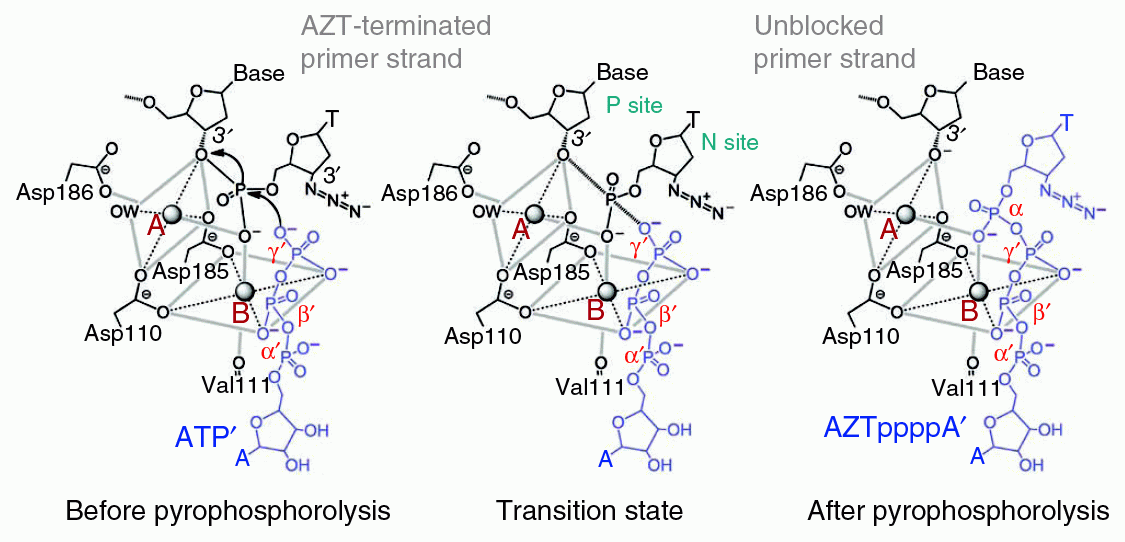
Proposed mechanism for ATP-mediated excision of AZT from DNA primer. The three Asp residues of HIV-RT which are involved are the same ones used for DNA-based RNA polymerization. Spheres A and B are Mg2+ ions, which are required for catalysis.
Submitted by: Marian Szebenyi, MacCHESS, Cornell University
