X-RAY RUNS: Apply for Beamtime
2017 Nov 1 - Dec 21
2018 Feb 7 - Apr 3
2018 Proposal/BTR deadline: 12/1/17
2018 Apr 11 - Jun 4
2018 Proposal/BTR deadline: 2/1/18
Thiamin (British spelling thiamine) is an important cofactor in many enzymatic reactions. Humans do not synthesize their own thiamin, which is why it is characterized as vitamin B1, an essential part of our diet. Most of the biochemical steps in the synthesis of thiamin by bacteria and plants are known, along with the enzymes that catalyze them. Although the protein TenI has been thought to be part of the thiamin pathway because it tends to cluster with other genes for thiamin synthesis and shares sequence similarity to some of them, a specific role in the pathway for TenI had never been identified. This question has now been addressed by a collaboration between Steven E. Ealick, the William T. Miller Professor in Cornell University Department of Chemistry and Chemical Biology, and his Cornell co-workers Ying Han, Abhishek Chatterjee and Yang Zhang, and Tadhg P. Begley, the Robert A. Welch Chair in Chemistry and D.H.R. Barton Professor of Chemistry at Texas A&M University and his Texas A&M co-workers Amrita Hazra and Rung-Yi Lai.
Using a variety of specially designed chemical assays and techniques, such as high pressure liquid chromatography and electrospray mass spectrometry, the team identified a biochemical role for the TenI enzyme of the soil bacterium Bacillus subtilis. It serves as a thiazole tautomerase, catalyzing the aromatization of (R,Z)-2-(2-carboxy-4-methylthiazol-5(2H)-ylidene)ethyl phosphate (cThz*-P) to 2-(2-carboxy-4-methylthiazol-5-yl)ethyl phosphate (cThz-P).
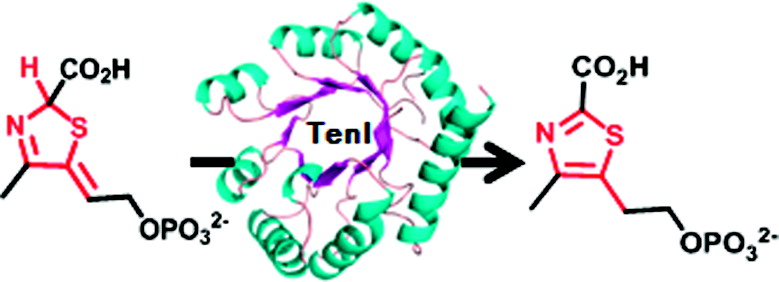
This rearrangement of the double bonds makes cThz-P a much better substrate than cThz*-P for the next step of the pathway, which greatly speeds up the synthesis of thiamin.
The researchers then grew crystals of TenI bound with the reaction product cThz-P, and brought them to CHESS beamline A1 to collect diffraction data. The structure of TenI alone had been solved a few years earlier by these same laboratories, and this earlier structure, available as Protein Data Bank entry 1YAD was used to help solve the structure of the enzyme product complex, which is now available as Protein Data Bank entry 3QH2.
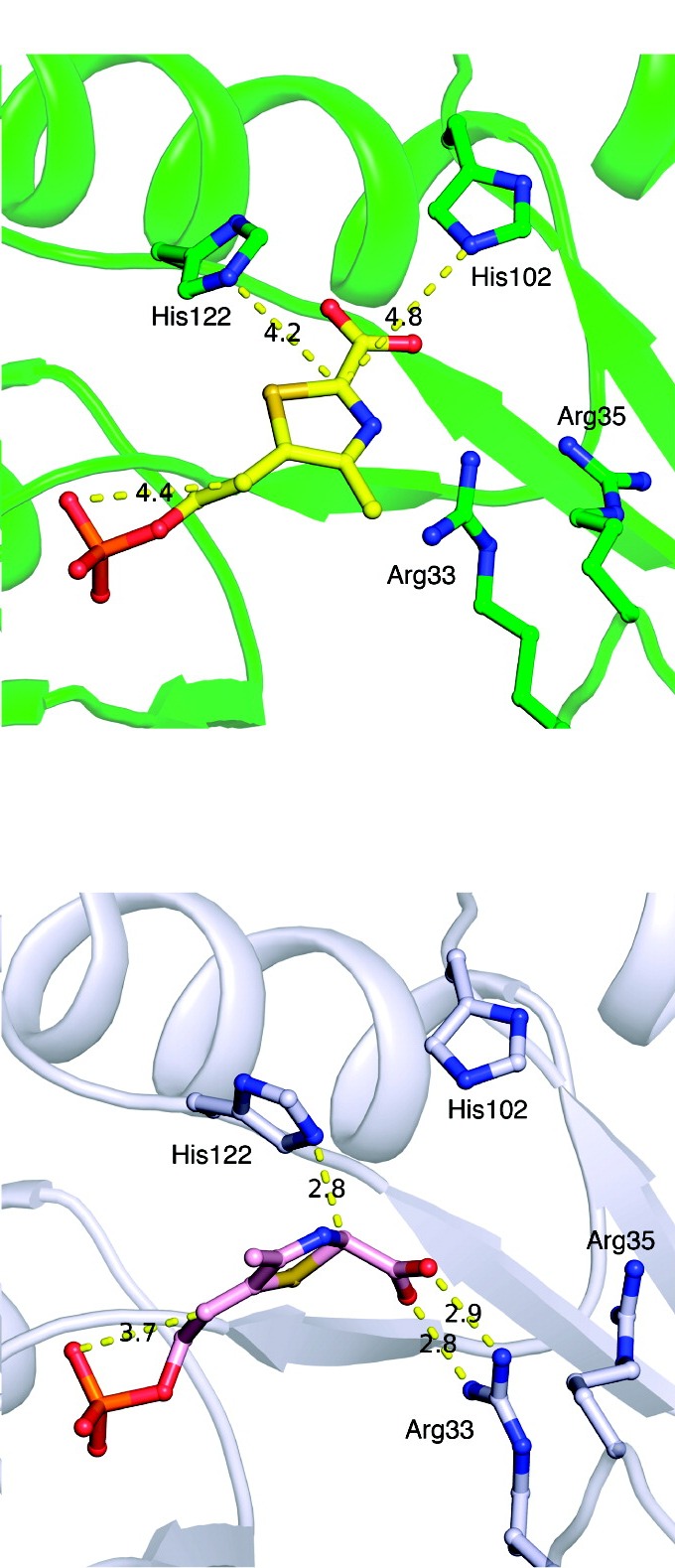
Using the coordinates of the complex, the researchers were able to identify the residues of TenI that are likely involved (Histidine 122, along with the phosphate of the substrate), and to propose a mechanism for the reaction. The identification of a function for TenI completes the identification of all of the enzymes needed for thiamin biosynthesis by the major bacterial pathway, thus improving our understanding of the biosynthesis of a vital nutrient.
A Missing Enzyme in Thiamin Thiazole Biosynthesis: Identification of TenI as a Thiazole Tautomerase
Amrita B. Hazra, Ying Han, Abhishek Chatterjee, Yang Zhang, Rung-Yi
Lai, Steven E. Ealick, and Tadhg P. Begley
J. Am. Chem. Soc., 133 (24), pp
9311-9319 (2011)
Submitted by: David J. Schuller, MacCHESS, Cornell University
