X-RAY RUNS: Apply for Beamtime
2017 Nov 1 - Dec 21
2018 Feb 7 - Apr 3
2018 Proposal/BTR deadline: 12/1/17
2018 Apr 11 - Jun 4
2018 Proposal/BTR deadline: 2/1/18
Eukaryotic cells, ranging from fungi to mammals to plants, are defined by their complex assortment of internal membrane-bound organelles, such as DNA-containing nuclei, energy-producing mitochondria, and many others. This allows them (us!) to keep important but incompatible activities separate, but it comes at a price: the proteins required for each organelle’s function, and the proteins destined for the outside of the cell (the plasma membrane), must be sorted to their separate destinations by a dedicated organelle, the Golgi apparatus. Failure to sort proteins into membrane vesicles targeted to the correct destinations leads to a broad spectrum of neurological and other disorders, including periventricular heterotopias with microcephaly, cranio-lenticulo-sutural dysplasia, and chylomicron retention disease (Olkkonen and Ikonen, 2006).
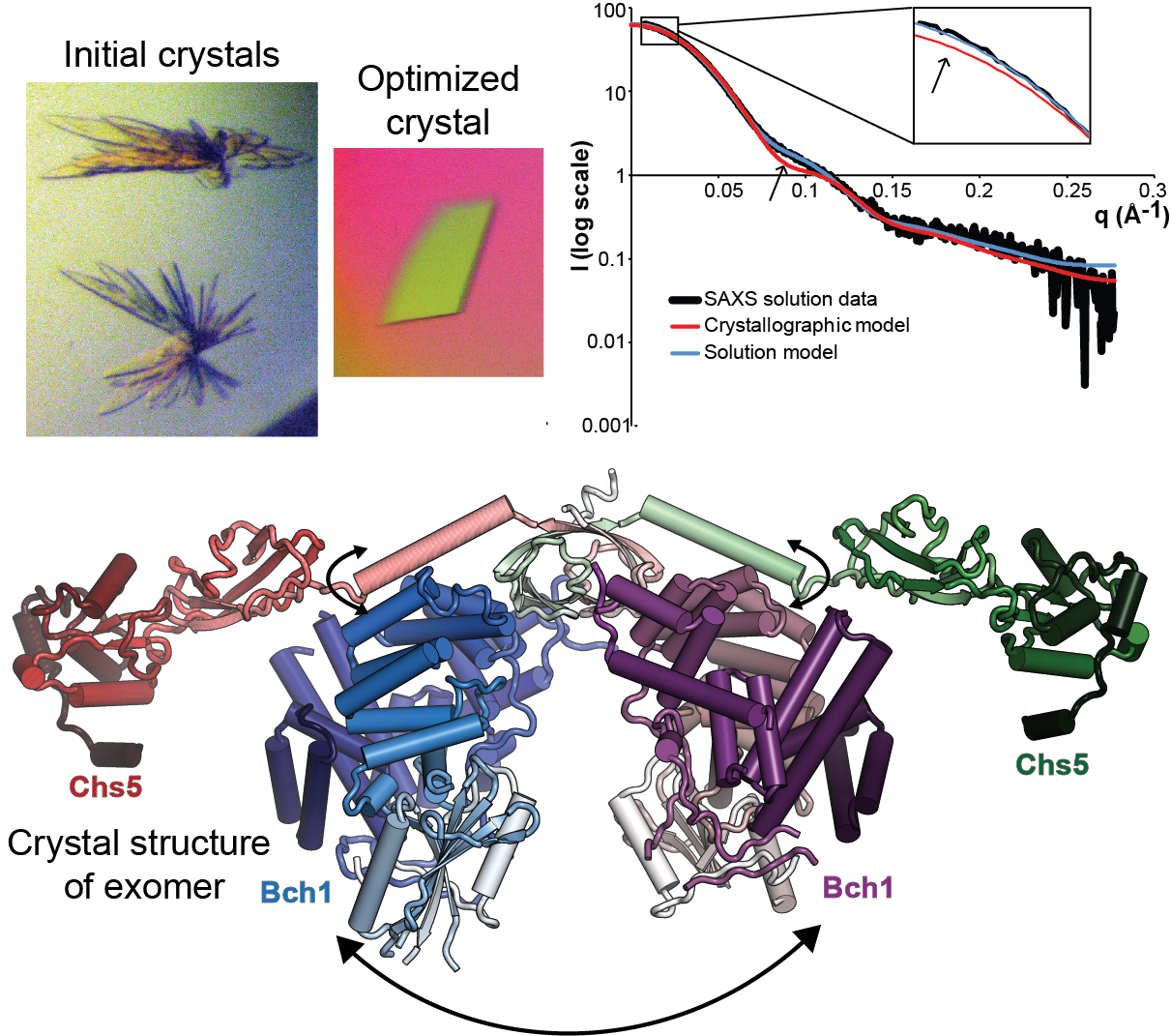
How, then, does the cell take a random mixture of membrane-bound proteins and sort then into their appropriate vesicle carriers, when the proteins are arranged more as a jumble on the floor than as an orderly assembly line, yet the vesicles are created by highly ordered protein cages? A new study conducted at CHESS beamlines A1 and F2 by postdoctoral associate Brian Richardson and assistant professor Chris Fromme of Cornell University, recently published in Structure, combines X-ray crystallography with small-angle X-ray scattering (SAXS) to identify an inherent structural flexibility in one such cargo sorting adaptor, exomer.
Exomer is a yeast cargo sorting adaptor that identifies a set of proteins requiring delivery to the plasma membrane at specific places and times in the cell cycle, and pulls them out of the general sorting pool into vesicles for regulated delivery. Prior studies suggested that exomer uses a modular architecture to identify multiple proteins, with four cargo-binding proteins interchangeably bound to a single core protein, Chs5 (Trautwein et al., 2006; Wang et al., 2006). This straightforwardly modular architecture makes exomer an excellent and very tractable model system for study of general cargo sorting across all eukaryotes. By solving the crystal structure of the core complex with one of the four cargo-binding proteins, Bch1, Richardson and Fromme showed that two copies of Bch1p are joined together by Chs5 at one end. Pairing this static crystal structure with SAXS data collected on the complex floating free in solution, they further showed that this arrangement allows exomer to open and close akin to a clamshell, with the tips likely at the membrane surface. Adding to this an earlier paper by Richardson and Fromme with graduate student Jon Paczkowski published in the EMBO Journal identifying a flexibly attached domain recruiting exomer to the membrane by way of the regulating protein Arf1, the group develops a model in which three pivots in exomer allow it to adapt to a flexible, diverse membrane environment.
References:
Olkkonen, V.M., and Ikonen, E. (2006). When intracellular logistics fails - genetic defects in membrane trafficking. J Cell Sci 119, 5031–5045.
Paczkowski, J.E., Richardson, B.C., Strassner, A.M., and Fromme, J.C. (2012). The exomer cargo adaptor structure reveals a novel GTPase-binding domain. The EMBO Journal 31, 4191–4203.
Trautwein, M., Schindler, C., Gauss, R., Dengjel, J., Hartmann, E., and Spang, A. (2006). Arf1p, Chs5p and the ChAPs are required for export of specialized cargo from the Golgi. EMBO J 25, 943–954.
Wang, C.-W., Hamamoto, S., Orci, L., and Schekman, R. (2006).
Exomer: a coat complex for transport of select membrane proteins from the trans-Golgi network to the plasma membrane in yeast. J. Cell Biol. 174, 973–983.
Submitted by: Richard Gillilan, MacCHESS, Cornell University
2/19/2013
