X-RAY RUNS: Apply for Beamtime
2017 Nov 1 - Dec 21
2018 Feb 7 - Apr 3
2018 Proposal/BTR deadline: 12/1/17
2018 Apr 11 - Jun 4
2018 Proposal/BTR deadline: 2/1/18
For some proteins, more is needed after their translation from DNA before they are functional. Small molecules are attached to certain amino acids in the protein in a process called post-translational modification. If the attachments are later removed, the behavior of the protein changes. The family of proteins called "sirtuins" is responsible for such removals in a number of biological systems. The sirtuins have attracted a great deal of interest from a broad biological community because of their suggested roles in the regulation of gene transcription and chromatin structure, as well as DNA repair and genome stability, cell metabolism, and perhaps most importantly, their potential connections to disease states (e.g. cancer) and aging. There are several members of the sirtuin family, and the majority of them catalyze the deacetylation of certain lysine amino acid residues which have been post-translationally modified by the addition of acetate. However, Sirt5, a sirtuin found in mitochondria, does not appear to work this way. Yeyun Zhou, a Cornell biophysics graduate student, set out to find out why Sirt5 is different, and what it does do instead of deacetylation.
To gain some insight into why Sirt5 was ineffective as a deacetylase, Yeyun determined the X-ray crystal structure for Sirt5 in complex with a thioacetyl peptide. It turned out that a buffer molecule, CHES, was also bound to Sirt5. This suggested to Yeyun and two of her mentors, Quan Hao, formerly the Director of MacCHESS and currently a professor at Hong Kong University, and Hening Lin, a professor in the Cornell Chemistry department whose laboratory focuses on the enzymology of sirtuin family members, that perhaps Sirt5 is optimally designed to bind compounds somewhat larger than acetate, and with an acyl group containing a negatively charged carboxylate (i.e. succinyl and malonyl moities). This conclusion was strongly supported by the determination of a crystal structure of a complex of Sirt5 with a succinyl peptide, which showed that the succinyl group fit very well into the active site. Indeed, the structural information obtained by Yeyun turned out to provide a powerful clue to the true enzymatic function of Sirt5, as the Lin group was able to show using an assay based on mass spectrometry that this protein, unlike previously characterized sirtuins such as Sirt1, was an effective catalyst for de-succinylation and de-malonylation, but not for deacetylation.
The research is reported in Science.
J. Du, Y. Zhou, X. Su, J.J. Yu, S. Khan, H. Jiang, J. Kim, J. Woo, J. H. Kim, B. H. Choi, B. He, W. Chen, S. Zhang, R. A. Cerione, J. Auwerx, Q. Hao,
and H. Lin; "Sirt5 is a NAD-dependent Protein Lysine Demalonylase and
Desuccinylase", Science 334, 806-809 (2011)
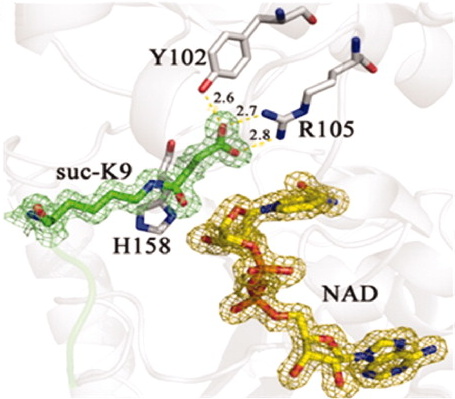
Close-up of succinyl peptide and NAD bound in Sirt5, showing interactions of succinyl with Tyr 102 and
Arg 105 of SirT5.
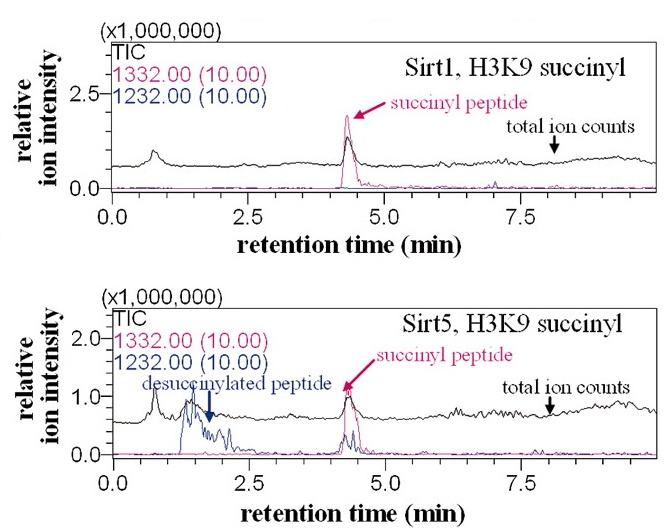
Sirt 5 (bottom) but not Sirt1 (top) removes a succinyl group from
a peptide. These chromatography-mass spectrometry traces show peaks
(pink) a the position of the succinylated peptide, but a peak
(blue) for the desuccinylated peptide appears only for Sirt5.
This is a beautiful first-hand example of how insights from X-ray crystallographic determinations of 3-D structures can significantly impact fundamentally important areas in biology and biomedicine. The work on Sirt5 carries some extremely important and exciting implications. One is that it points to a brand new class of post-translational protein modifications in biology (namely succinylation and malonylation). Perhaps equally important, the work catalyzed the collaboration between Rick Cerione’s laboratory (Cerione is the PI of MacCHESS) in Molecular Medicine at Cornell, and the Lin laboratory, that has shown that the enzymatic activity of Sirt5 is essential for the transformed phenotypes exhibited by various cancer cell lines. Indeed, Cerione, Lin and their colleagues now suspect that Sirt5 and its effects on various metabolic enzymes play a key role in cancer metabolism, which has led to their recently being awarded a Transformative Grant from the National Cancer Institute to pursue this hypothesis. Thus, from the structural determinations performed by a young student at CHESS has emerged an exciting new area in cancer research, further demonstrating the critical roles played by structural determinations in biomedicine.
Submitted by: Marian Szebenyi, MacCHESS, Cornell University
01/31/2012
